From Variant to Samples
In Mosaic, there are several ways to get sample information for a variant.
Genotype columns in variants table
The quickest way to get sample information for a variant is to look in one of two genotype columns that are available in the variants table. The genotype count column shows you the number of hets and homs that have that variant.

Additionally, there is the genotype column which shows the genotypes of the samples using graphical icons. An empty circle is for REF/Unknown, a half circle for Hets, and a filled-in circle for Homs. In projects with too many samples to show at once, you can click on down arrow next to the Genotype header to select the samples to show.

These two columns are great for quickly getting sample genotype information when scrolling through multiple variants, however if you want to look more closely at the samples for one or more variants the next two options may be more useful.
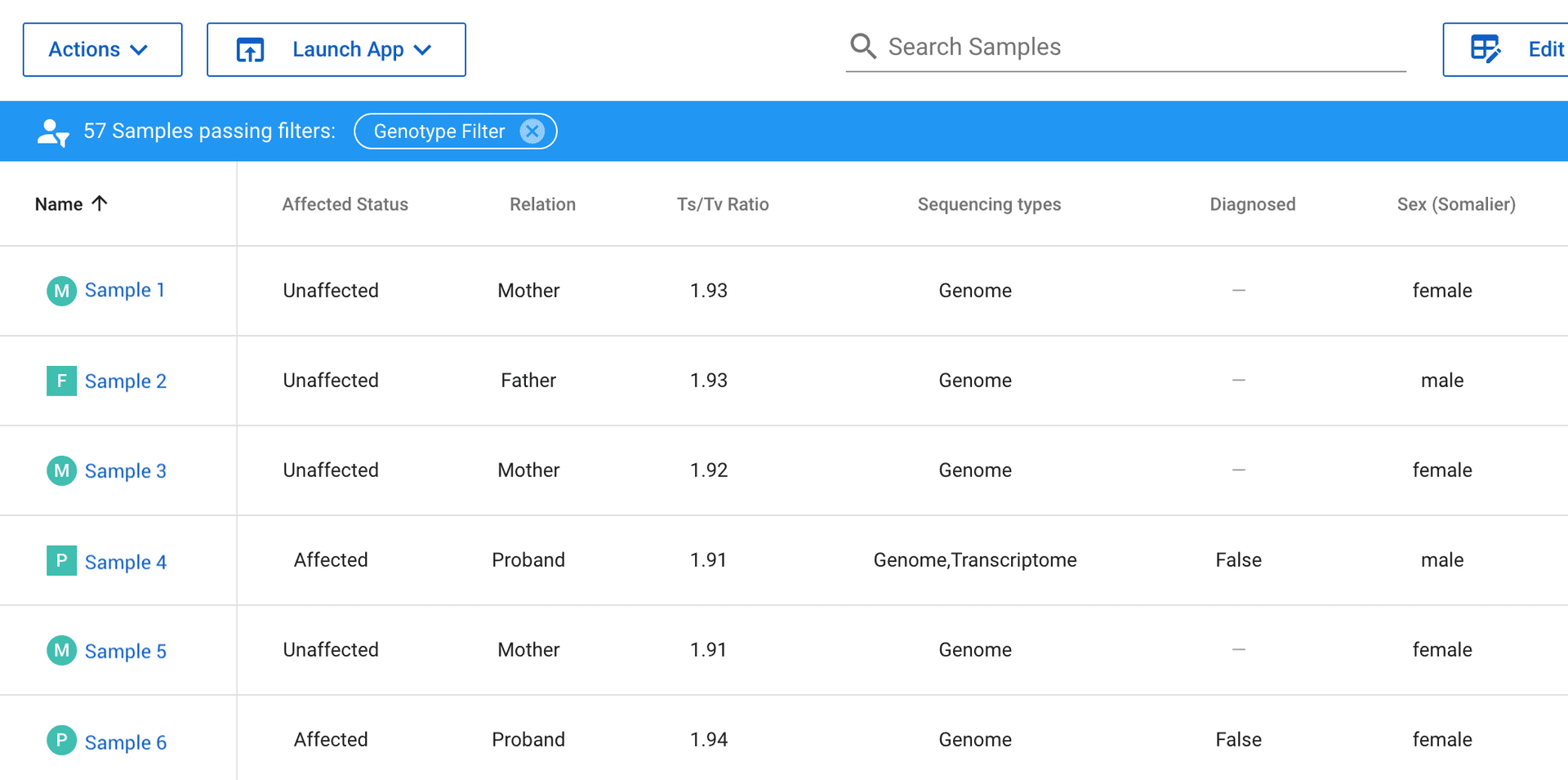
Samples table view
To look more at the samples more in depth and see the attribute values, going to the samples table is very useful. You can do this by selecting one or more variants in the variants table, going to the Actions button, and clicking 'View samples with these variants'. This will show you the passing samples in the samples table. You can then customize the samples table to show any sample attributes as columns. This is very useful for quickly scrolling through all the samples and scanning the values of all the attributes of interest.
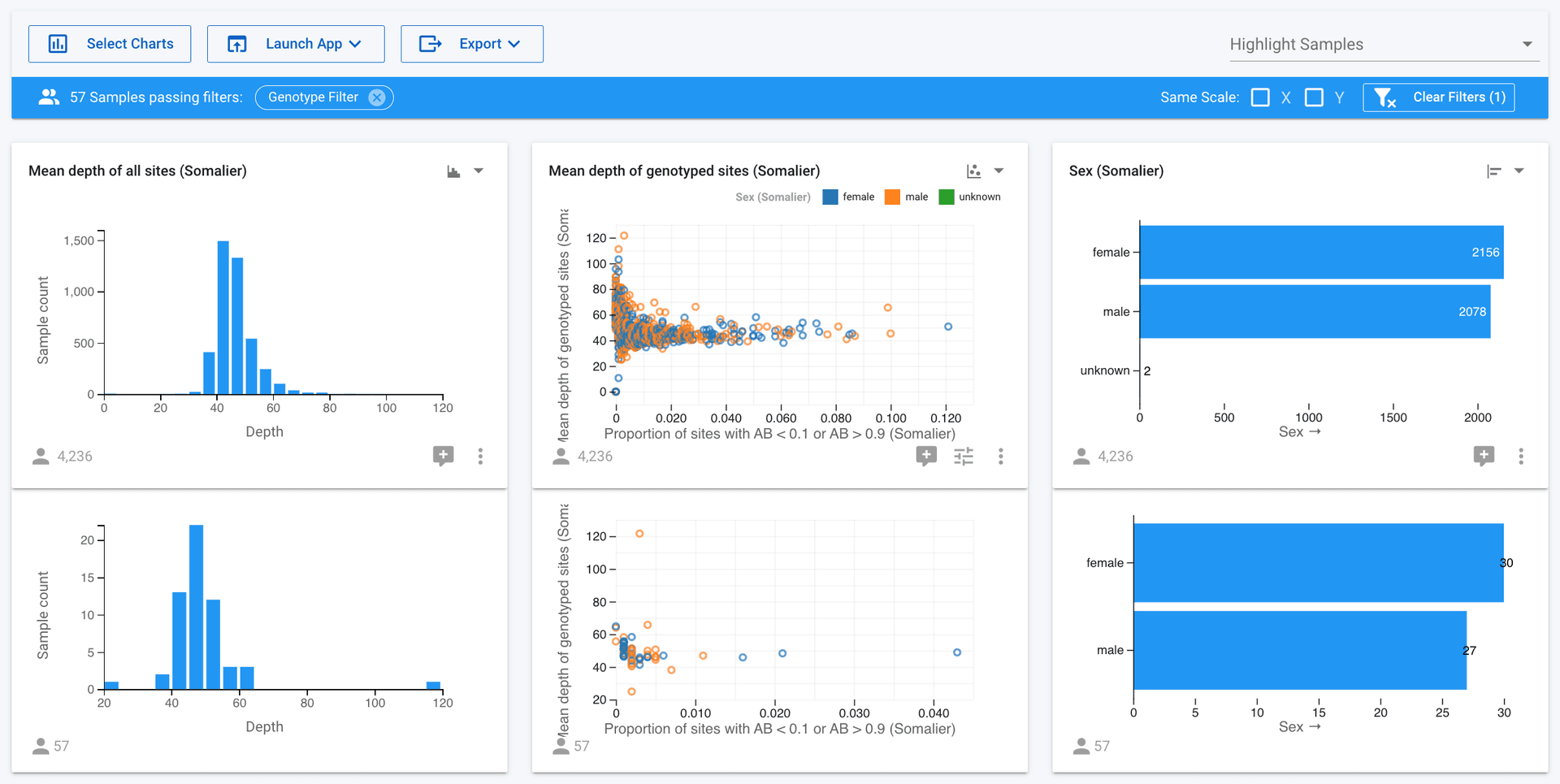
Samples analytics view
Additionally, you can see the samples that have a particular variant(s) as charts in analytics. To do this select one or more variants in the variants table, go to the Actions button, and click 'View samples with these variants in Analytics'. You can now see how the selected samples compare against the rest of the samples in the project. Additionally, you can further filter by attributes to dig deeper into the data. This is very useful for exploring the data to answer questions or identify things of interest.