How Mosaic Offers Free Unlimited Genomic Samples
Mosaic allows users to add as much genomic data (CRAM, BAM, VCF) as they want to their Mosaic projects. This is true for paid accounts but surprisingly this is also true for free accounts (if you haven't already you can sign up and go crazy right now). As far as I'm aware, we are the only genomic cloud platform that offers this or perhaps more accurately can offer this. We are able to do this (without going bankrupt) due to how Mosaic is designed, which lets us operate very differently than other genomic cloud platforms.
Most genomic cloud platforms are centralized and require a user to upload all their genomic data to a vendor's cloud. This is a straightforward solution, but it also means that the first step for a new user is doing something prohibitively costly to offer for free.
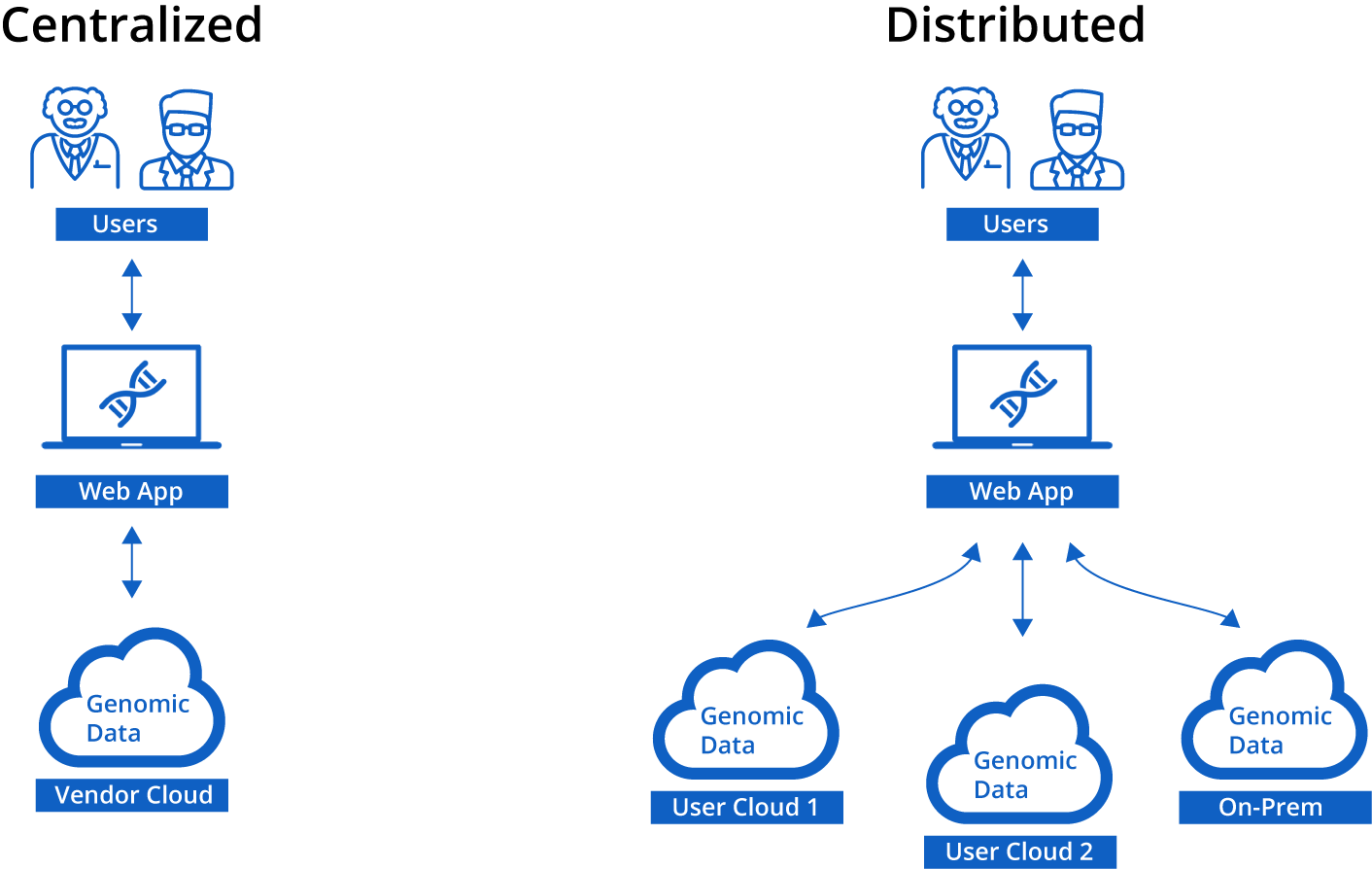
In contrast, Mosaic was designed to be a distributed system that doesn't impose restrictions on the underlying storage but rather works as a lightweight layer on top of storage. This allows Mosaic to integrate data from different clouds and on-prem systems and present them as a single, simple UI. But more relevant to this post, this design also means that users can add genomic data to Mosaic that is already in the cloud on their existing S3 buckets without moving them. This is the trick. Since we are not copying the data to our own S3 buckets, there is no extra cost and we can add huge numbers of samples to Mosaic for relatively cheap. For any organization that already has data sitting on their own S3 buckets (which are many), then this is a great way to get much more out of that resource.
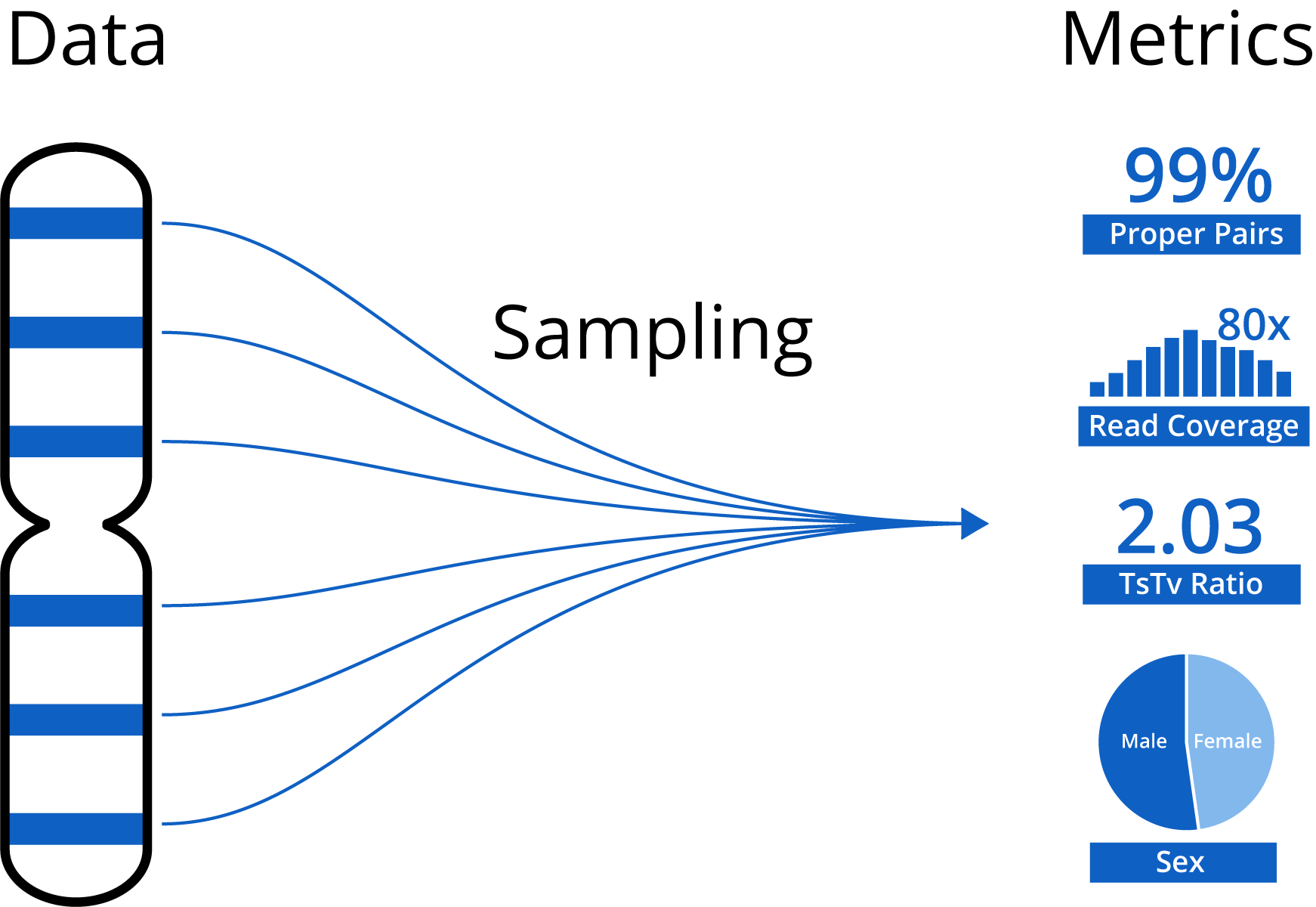
However, Mosaic is a lot more than just project management. We also automatically generate a host of statistics and QC metrics, which (along with user-supplied phenotypes) are visualized and interacted with via our analytics functionality. Generating the statistics and metrics with standard methods takes several hours per sample and is too costly to offer for free, but to get the full value of Mosaic it's essential we still generate these statistics for free accounts. To accomplish this, we use the sampling methodology that we developed over years of working on real-time sampling of genomic data as part of the iobio project. Instead of processing the entire multi-gb file, we process a small subset of the data to generate global statistics. We've shown that you only need a surprisingly small amount of the data to generate very accurate results (bam.iobio publication). Using these iobio tricks, we can generate these statistics and metrics in near real-time and at basically no additional cost.
Due to Mosaic's distributed design and use of real-time sampling, we are able to offer the bulk of Mosaic functionality for free for all users without any limitations on the number of samples. We think this is unique among commercial cloud genomic platforms and we are really proud and excited about this accomplishment. We hope that it becomes a valuable resource for collaboration in the genomics community.
-- Chase
Here at Frameshift we are hard at work improving how people interact with genomic data and are actively applying these ideas into our flagship product Mosaic. If you are interested in genomic visualization and genome-scale analytics (or even just like to talk about these ideas) get in touch info@frameshift.io!
OR
Create a free account in Mosaic today.